Translation of NK Cell CAR Therapy to the Clinic: Critical Role of Performance & Clinical-scalability
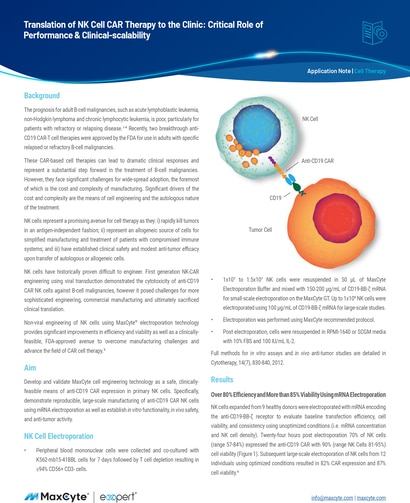
The prognosis for adult B-cell malignancies, such as acute lymphoblastic leukemia (ALL),1,2 non-Hodgkin lymphoma (NHL)3 and chronic lymphocytic leukemia (CLL),4 is poor, particularly for patients with refractory or relapsing disease. Recently, two breakthrough anti-CD19 CAR-T cell therapies were approved by the FDA for use in adults with specific relapsed or refractory B-cell malignancies.
Read MoreOptimizing CRISPR/Cas-Mediated CAR Knockin in Primary Human T Cells Using MaxCyte Electroporation
MaxCyte electroporation optimized CAR knockin to enable stable expression of tumor-targeting receptors in human T cells, while maintaining high cell viabilities and functionality.
Read MoreOptimizing Transposon-Based Gene Delivery for Cell Therapy Applications with the MaxCyte® ExPERT Scalable Electroporation Platform
MaxCyte electroporation efficiently engineered primary human T cells with piggyBac transposons to express CARs, while maintaining high cell viabilities, cell yields and functionality.
Read MoreMaxCyte Electroporation Enables the Production of Assay-Ready Cells Suitable for Ion Channel Studies
In this study, we show scalable, efficient transfection in a variety of cell lines and stem cells and present case studies demonstrating the suitability of assay-ready cells as an alternative to stable cell lines for ion channel research, assay development and drug discovery.
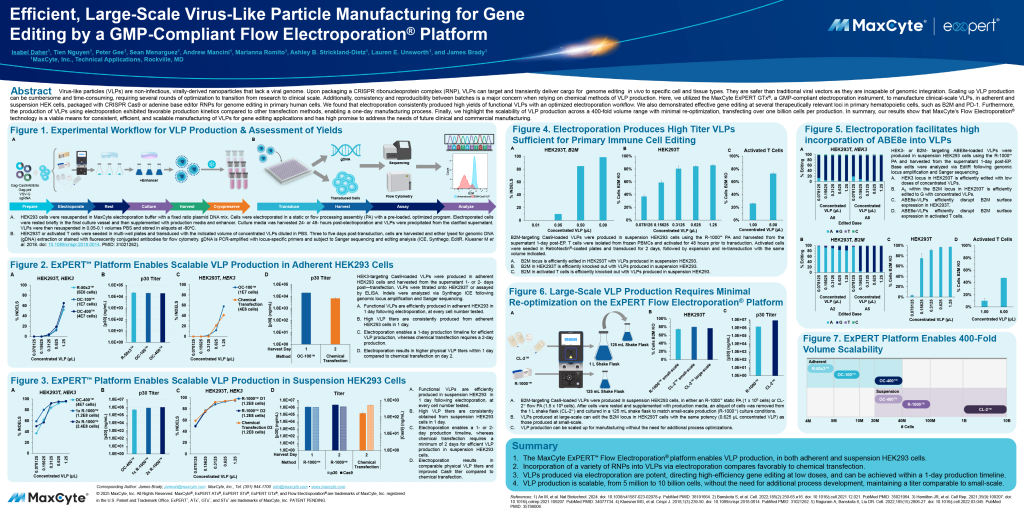
Read MoreEfficient, Large-Scale Virus-Like Particle Manufacturing for Gene Editing by a GMP-Compliant Flow Electroporation Platform
The ExPERT GTx® efficiently manufactured clinical-scale VLPs in adherent and suspension HEK cells, packaged with CRISPR-Cas9 or adenine base editor RNPs, for genome editing in primary human cells.
Read MoreHighly Efficient Engineering of Difficult-to-Transfect Immune Cells Using MaxCyte® Electroporation
MaxCyte enabled transient and stable expression of CARs/TCRs in T cells, NK cells and macrophages through high-efficiency transfection of mRNA, DNA encoding transposons/transposases, or CRISPR ribonucleoproteins (RNPs) and homology-directed repair (HDR) templates into these hard-to-transfect cells
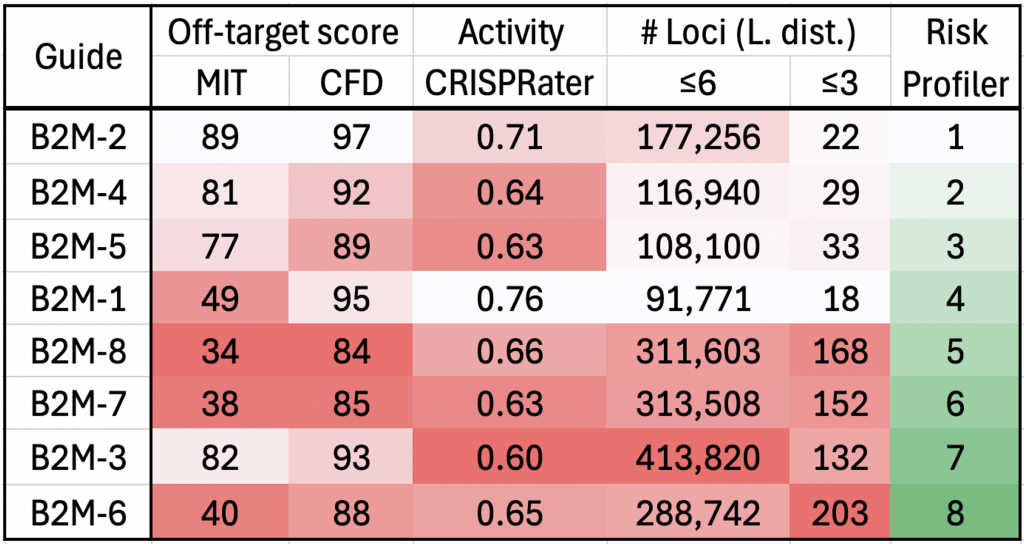
Read MoreGuide Profiler™: A Genetic Variant-Aware Computational Tool for Improved Guide RNA Selection for CRISPR-Based Therapeutic Applications
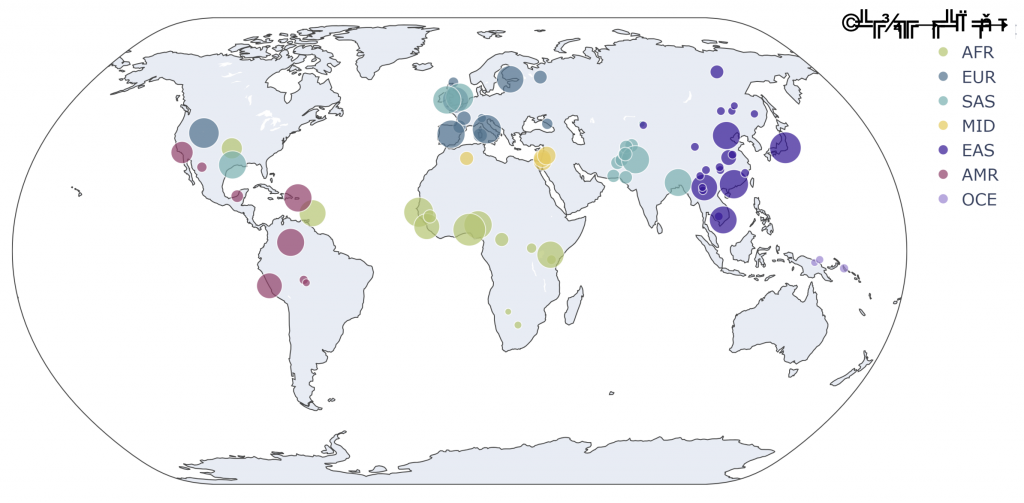
Guide Profiler evaluates off-target risk during the guide RNA (gRNA) selection process. Using in silico off-target enumeration and computational methods, the tool rapidly screens large numbers of gRNA sequences and makes an informed list of viable candidates. Guide Profiler will allow therapy makers to take into account ancestry of target donor and intent-to-treat populations, as well as to assess preparedness for diverse populations in clinical trial design.
Read MoreSAFER Detection™ for Efficient Interrogation of DNA Rearrangements in Gene-Edited Human Cells
In this study, we introduce SAFER Detection (Selective Amplification for Efficient Rearrangement Detection), a tagmentation and next-generation sequencing method designed to enable quantitative detection of chromosomal rearrangement breakpoints with single base resolution. The method is capable of classifying rearrangements resulting from on-target and off-target editing by programmable nucleases.
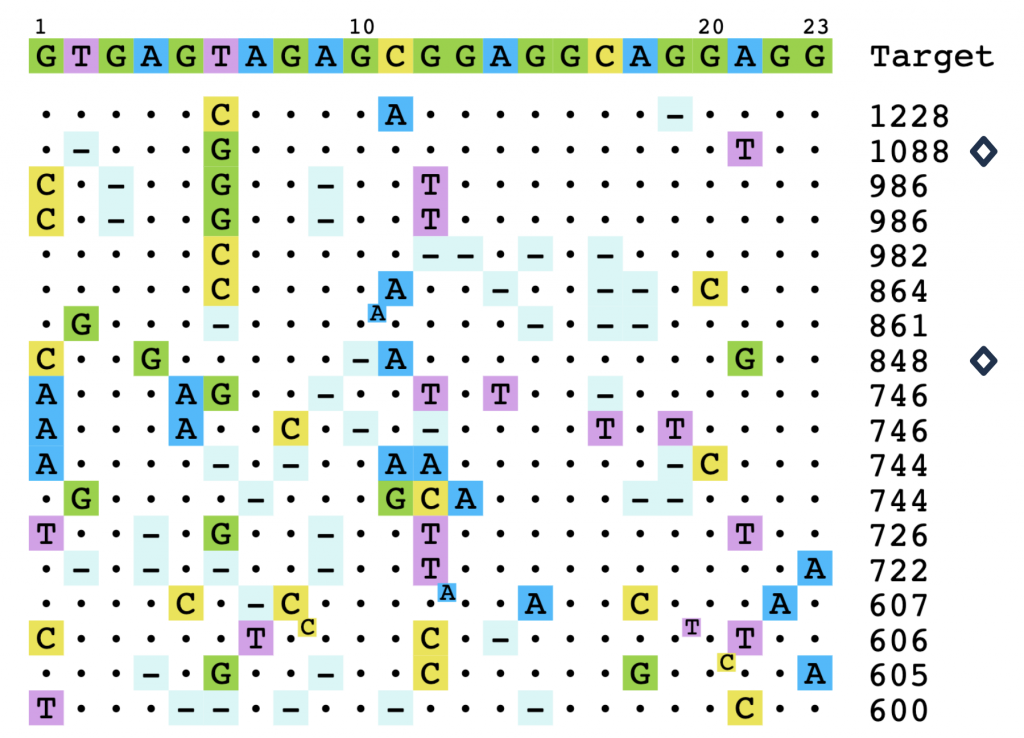
Read MoreVariant-Aware Off-Target Analysis for Informed Guide RNA Selection
Here, we generated data for guides targeting the PCSK9 gene using the CRISPR-Cas9 nuclease system, while considering common variants in major human superpopulations. SeQure DX™ assays enabled a multistep approach that reduced off-target risk in the guide RNA selection process.
Read MoreONE-seq™ for Variant-Aware Therapeutic Guide Selection
ONE-seq leverages computational tools and biochemical assays to nominate candidate off-target sites across thousands of genomes with high sensitivity. Here, we present an application of ONE-seq to identify guides with the lowest potential off-target editing risk.
Read More